| Name |
heterogeneous nuclear ribonucleoprotein U |
interleukin 7 receptor |
| Structure |
No pdb structure
|
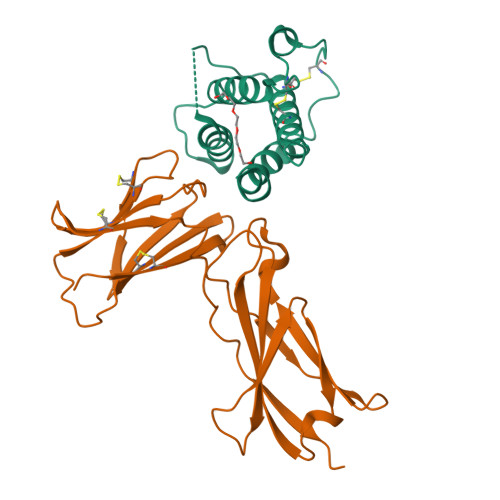
|
| GO Annotations |
Cellular Component |
|
|
| Molecular Function |
|
|
| Biological Process |
|
|
| Pathways |
|
|
| Diseases |
None
|
|
| GWAS |
No GWAS studies recorded
|
|
| Interacting Genes |
42 interacting genes:
ACTB
BTRC
CASP3
CD5
CDKN2A
CEBPA
CR2
DUX4
ELL
EP300
ERG
GRIN1
GRIN2D
GTF2H1
HMGB1
HNRNPH3
HSPB1
IL7R
KAT2B
LINC00624
NDN
NDRG1
NEDD4
NR3C1
NRON
PIN1
PLK1
POU3F4
PRMT1
PRPF40A
PTPN11
RBPMS2
SMN1
SREK1
STAU1
SUMO2
SYK
TCERG1
UBE2I
WBP4
YAP1
ZNF689
|
110 interacting genes:
AGTRAP
ALYREF
APOL3
CIRBP
CPSF1
CRLF2
DDX21
DDX39B
DDX3X
DDX5
DHX36
DHX9
EIF2AK2
ELAVL1
EMG1
FAU
FUS
FYN
G3BP1
H1-10
H1-2
H1-4
H2BC21
HNRNPA0
HNRNPA3
HNRNPAB
HNRNPC
HNRNPD
HNRNPDL
HNRNPH3
HNRNPL
HNRNPR
HNRNPU
HNRNPUL1
HNRNPUL2
IL2RG
IL7
ILF2
ILF3
JAK1
JAK3
KIT
LYN
MALL
MAP4
MS4A1
NCL
NONO
PABPC1
PABPC4
PABPN1
PIK3R1
PTBP1
PTK2B
PTMA
PURA
PURB
QKI
RACK1
RAD21
RBM3
RBMX
RPL15
RPL18
RPL22
RPL29
RPL30
RPL31
RPL6
RPL7
RPL8
RPS20
RPS3
RPSA
RRAGA
RSL1D1
SAFB
SDC4
SF1
SF3A1
SF3B1
SNRNP70
SNRPA
SNRPB
SNRPD1
SNRPD2
SNRPD3
SNRPE
SNRPF
SNRPG
SRP14
SRP9
SRSF3
SRSF9
SSB
STAT3
STAT5A
STAT5B
SYNCRIP
TMEM120B
TOE1
TOP1
TSLP
U2AF1
U2AF2
YBX1
YBX3
YWHAE
YWHAG
ZNF787
|
| Entrez ID |
3192 |
3575 |
| HPRD ID |
04185
|
00893
|
| Ensembl ID |
ENSG00000153187
|
ENSG00000168685
|
| Uniprot IDs |
Q00839
Q96BA7
|
P16871
|
| PDB IDs |
Not available
|
3DI2
3DI3
3UP1
5J11
6P50
6P67
7OPB
|
| GO Terms Enriched among Interactors |
|
|