| Name |
glutamate ionotropic receptor AMPA type subunit 2 |
general transcription factor IIIC subunit 2 |
| Structure |
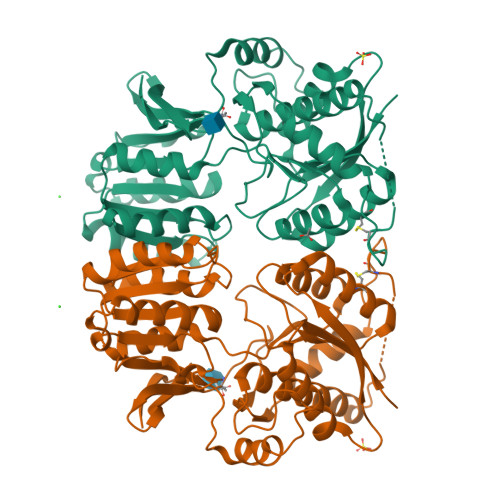
|
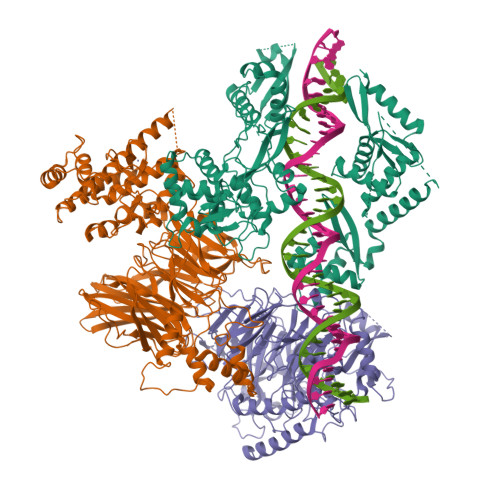
|
| GO Annotations |
Cellular Component |
|
|
| Molecular Function |
|
|
| Biological Process |
|
|
| Pathways |
|
|
| Drugs |
- Glutamic acid
- Butabarbital
- Butalbital
- Talbutal
- Pentobarbital
- Secobarbital
- Metharbital
- Thiopental
- Primidone
- Methylphenobarbital
- Ethanol
- Phenobarbital
- Quinidine barbiturate
- Amobarbital
- Aprobarbital
- Butobarbital
- Heptabarbital
- Hexobarbital
- Barbital
- Dihydro-2-thioxo-5-((5-(2-(trifluoromethyl)phenyl)-2-furanyl)methyl)-4,6(1H,5H)-pyrimidinedione
- (S)-DES-ME-AMPA
- (S)-AMPA
- 2-Amino-3-(5-Tert-Butyl-3-(Phosphonomethoxy)-4-Isoxazolyl)Propionic Acid
- Iodo-Willardiine
- Fluoro-Willardiine
- Quisqualic acid
- (S)-2-Amino-3-(1,3,5,7-Pentahydro-2,4-Dioxo-Cyclopenta[E]Pyrimidin-1-Yl) Proionic Acid
- (2S)-2-Ammonio-3-[5-(2-methyl-2-propanyl)-3-oxido-1,2-oxazol-4-yl]propanoate
- FG-9041
- Bromo-Willardiine
- Willardiine
- 2-Amino-3-(3-Hydroxy-7,8-Dihydro-6h-Cyclohepta[D]-4-Isoxazolyl)Propionic Acid
- Aniracetam
- THIO-ATPA
- Talampanel
- CX-717
- N,N'-[biphenyl-4,4'-diyldi(2R)propane-2,1-diyl]dimethanesulfonamide
- 2,3,6A,7,8,9-HEXAHYDRO-11H-[1,4]DIOXINO[2,3-G]PYRROLO[2,1-B][1,3]BENZOXAZIN-11-ONE
- (3S)-3-cyclopentyl-6-methyl-7-[(4-methylpiperazin-1-yl)sulfonyl]-3,4-dihydro-2H-1,2,4-benzothiadiazine 1,1-dioxide
- (3R)-3-cyclopentyl-7-[(4-methylpiperazin-1-yl)sulfonyl]-3,4-dihydro-2H-1,2-benzothiazine 1,1-dioxide
- (3R)-3-cyclopentyl-6-methyl-7-[(4-methylpiperazin-1-yl)sulfonyl]-3,4-dihydro-2H-1,2-benzothiazine 1,1-dioxide
- Fluciclovine (18F)
|
None
|
| GWAS |
- Triglyceride levels x loop diuretics use interaction ( 31806883
)
|
|
| Interacting Genes |
21 interacting genes:
CTTN
CYLD
DRD2
FXR1
GAPDH
GRIA3
GRIP2
GSK3B
GTF3C2
ITGB3
NSF
PICK1
PRKCA
RAC1
RNF167
SDCBP
SPTAN1
SPTBN1
SQSTM1
SYT3
TSPAN7
|
10 interacting genes:
GRIA2
GTF3A
GTF3C4
MECOM
PTEN
RB1
SUMO2
TBP
VAC14
VHL
|
| Entrez ID |
2891 |
2976 |
| HPRD ID |
04726
|
05347
|
| Ensembl ID |
ENSG00000120251
|
ENSG00000115207
|
| Uniprot IDs |
A0A994J4F1
P42262
|
Q8WUA4
|
| PDB IDs |
2WJW
2WJX
2XHD
3R7X
3RN8
3RNN
3UA8
5H8S
5YBF
5YBG
5ZG0
5ZG1
5ZG2
5ZG3
7F3O
8I0B
|
8CLI
8CLJ
8CLL
|
| GO Terms Enriched among Interactors |
|
- Transcription By RNA Polymerase III
- PML Body
- RNA Polymerase III General Transcription Initiation Factor Activity
- Positive Regulation Of RNA Biosynthetic Process
- Positive Regulation Of DNA-templated Transcription
- DNA-templated Transcription
- RRNA Transcription
- Enzyme Binding
- DNA-binding Transcription Factor Binding
- Positive Regulation Of RNA Metabolic Process
- Nucleoplasm
- Positive Regulation Of Nucleobase-containing Compound Metabolic Process
- Positive Regulation Of Transcription By RNA Polymerase II
- Negative Regulation Of Synaptic Vesicle Clustering
- PAS Complex
- Negative Regulation Of G1/S Transition Of Mitotic Cell Cycle
- Negative Regulation Of Cell Cycle G1/S Phase Transition
- Regulation Of Transcription By RNA Polymerase II
- Nucleobase-containing Compound Biosynthetic Process
- Phosphatidylinositol-3,4-bisphosphate 3-phosphatase Activity
- Negative Regulation Of Keratinocyte Migration
- Inositol-1,3,4,5-tetrakisphosphate 3-phosphatase Activity
- Inositol-1,3,4,5,6-pentakisphosphate 3-phosphatase Activity
- DNA-templated Transcription Initiation
- Positive Regulation Of Macromolecule Biosynthetic Process
- Positive Regulation Of Biosynthetic Process
- Ligand-gated Monoatomic Cation Channel Activity
- Positive Regulation Of Ubiquitin-dependent Protein Catabolic Process
- Phosphatidylinositol Biosynthetic Process
- Central Nervous System Myelin Maintenance
- Negative Regulation Of Protein Phosphorylation
- Rhythmic Synaptic Transmission
- Chromatin Lock Complex
- Positive Regulation Of Macromolecule Metabolic Process
- Regulation Of DNA-templated Transcription
- Regulation Of RNA Biosynthetic Process
- DNA Binding
- Histone H3K9 Monomethyltransferase Activity
- Dendritic Spine
- AMPA Glutamate Receptor Activity
- Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase Activity
- Positive Regulation Of Proteolysis Involved In Protein Catabolic Process
- Regulation Of G1/S Transition Of Mitotic Cell Cycle
- Myelin Sheath Adaxonal Region
- Negative Regulation Of Nervous System Development
- Negative Regulation Of Mitotic Cell Cycle Phase Transition
- Negative Regulation Of Peptidyl-serine Phosphorylation
- Regulation Of Synaptic Vesicle Clustering
- Negative Regulation Of Wound Healing, Spreading Of Epidermal Cells
- Phosphatidylinositol Phosphate Phosphatase Activity
|