| Name |
PDZ binding kinase |
G protein pathway suppressor 2 |
| Structure |
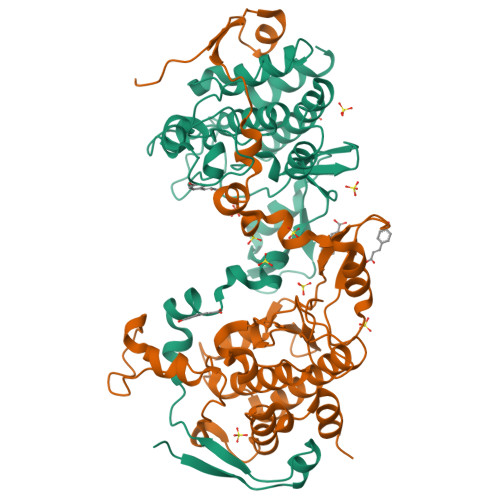
|
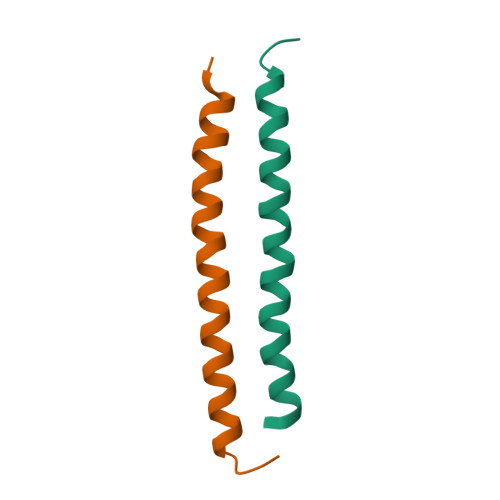
|
| GO Annotations |
Cellular Component |
|
|
| Molecular Function |
|
|
| Biological Process |
|
|
| Pathways |
None
|
|
| GWAS |
No GWAS studies recorded
|
- Liver enzyme levels (alkaline phosphatase) ( 33972514
)
|
| Interacting Genes |
25 interacting genes:
ARAF
CCNB1
CDK1
CEP126
DLG1
GPRASP1
GPS2
H2AX
LNX1
MCRS1
NSFL1C
PPP1CA
PRC1
PTEN
PTOV1
RAF1
SMAD4
TERT
TP53
TRIM37
UBC
VCP
ZBTB26
ZFP64
ZNF526
|
53 interacting genes:
ACTMAP
AKAP8L
ATF4
ATF5
BAG4
BRME1
CCNA1
CHD3
CNOT2
CYSRT1
DAZAP2
EP300
FAM168B
FHL5
GOLGA2
HDAC1
HDAC3
HNRNPH1
HOXA1
INTS11
KRT27
KRT31
KRT34
KRT36
KRTAP11-1
KRTAP13-2
KRTAP3-1
KRTAP3-3
KRTAP6-1
KRTAP6-2
KRTAP6-3
MAP3K7CL
NCOR1
NDOR1
NR0B2
OIP5
PBK
POU2AF1
PRMT6
PRR22
RBPMS
SESTD1
SETDB1
SMUG1
SPDL1
TBL1X
TBL1XR1
TFIP11
TP53
TP53BP2
TRIP6
UBTD2
VPS37C
|
| Entrez ID |
55872 |
2874 |
| HPRD ID |
17822
|
11878
|
| Ensembl ID |
ENSG00000168078
|
ENSG00000132522
|
| Uniprot IDs |
Q96KB5
V9HWH0
|
Q13227
|
| PDB IDs |
5J0A
|
2L5G
|
| GO Terms Enriched among Interactors |
|
|